Jul
23
2021
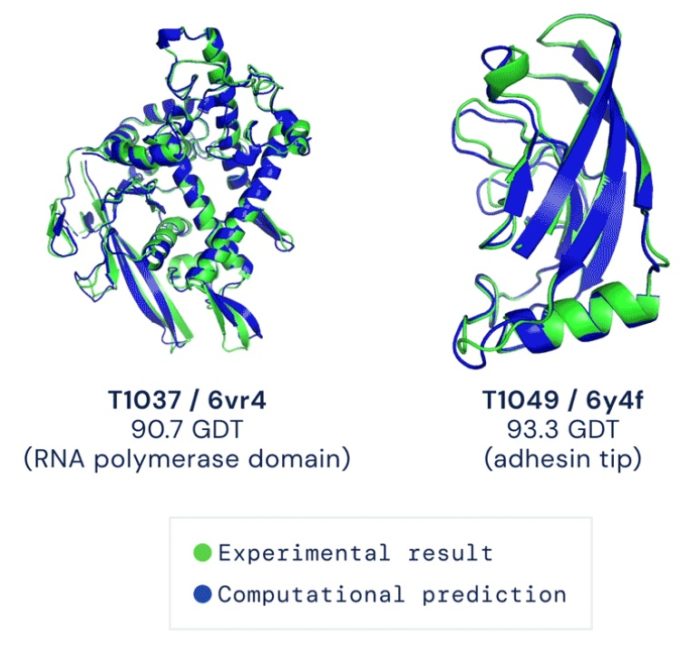 In 2003 the largest ever international cooperative scientific project was completed, at a cost of about $1 billion – the mapping of the human genome. This came with much fanfare, with the media hyping all the medical benefits that would soon flow. Of course, basic science progress often precedes clinical applications by decades, so the hype was not necessarily wrong, just premature. But it was an immediate boon to research, and those benefits are being felt today.
In 2003 the largest ever international cooperative scientific project was completed, at a cost of about $1 billion – the mapping of the human genome. This came with much fanfare, with the media hyping all the medical benefits that would soon flow. Of course, basic science progress often precedes clinical applications by decades, so the hype was not necessarily wrong, just premature. But it was an immediate boon to research, and those benefits are being felt today.
Perhaps the next big mapping project in biology is the human proteome, the characterization of every human protein. (I’ll also give a nod to the connectome project, the mapping of every connection in the human brain, but that will likely take much longer.) A new study published in Nature announces a significant leap forward in mapping the human proteome, using artificial intelligence (AI), specifically AlphaFold2 developed by DeepMind. To understand what they accomplished, however, we need to go over some basic concepts and terminology.
A gene is essentially a code for a sequence of amino acids, which make up proteins. So if we have mapped the entire sequence of bases (of which there are four – GATC) in a gene, we know the sequence of amino acids in the protein it codes for. So then, you might ask, if we have already mapped all the human genes, why is that not the same thing as a map of all the human proteins? This is because a protein is more than just a sequence of amino acids. A short chain of 2 or more amino acids is called a peptide, and a long chain is therefore a polypeptide. But we still don’t have a protein. A protein is a polypeptide that folds itself into a specific three-dimensional structure. It is that three-dimensional structure which determined the function and properties of the protein.
Continue Reading »
Dec
01
2020
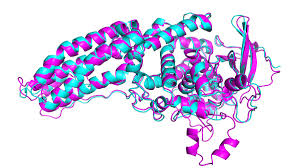 I wasn’t planning on writing about artificial intelligence (AI) two days in a row, but I also can’t plan the news. I also couldn’t pass up this item – London-based AI company, DeepMind, has mostly solved the extremely difficult problem of protein folding. If you are not already familiar with this issue, this may not sound like a big deal, but it is. So first lets’ give some background on the problem itself.
I wasn’t planning on writing about artificial intelligence (AI) two days in a row, but I also can’t plan the news. I also couldn’t pass up this item – London-based AI company, DeepMind, has mostly solved the extremely difficult problem of protein folding. If you are not already familiar with this issue, this may not sound like a big deal, but it is. So first lets’ give some background on the problem itself.
Biology is largely about proteins. Proteins are what genes code for, they make up enzymes, receptors, structural building blocks, antibodies, the basic machinery of cells, and more. Yes, lipids and carbohydrates are critical as well, but these are largely chaperoned by proteins. Proteins determine whether a cell is a liver or heart cell, and largely whether an organism is a human or sea cucumber.
Proteins are comprised of a sequence of amino acids, from a repertoire of 20 different amino acids. The specific sequence of amino acids is what is determined by the GATC genetic code in DNA, with three-letter codes for each amino acid. But a protein is more than just a sequence of amino acids. By itself a long chain of amino acids is a polypeptide – it doesn’t become a protein until that long chain is folded into a unique three-dimensional shape. Predicting how a long chain of amino acids will fold into a precise shape is the protein folding problem.
This is more difficult than it may at first seem. Imagine a chain with each link one of 20 possible different shapes, and that chain can be hundreds or even thousands of links long (titin is the largest known protein; its human variant consists of 34,351 amino acids). The number of possible ways to fold the protein gets magnified with each additional link. The resulting possibilities is staggering – too much for even the most powerful computer to crunch through. Determining how a sequence of amino acids actually folds is therefore determined mostly by direct laboratory study, using techniques such as X-ray crystallography and NMR spectroscopy. But this takes a long time – years for the largest proteins.
Continue Reading »
Sep
19
2011
The rather provocative title of this post refers to the at home science game, Foldit. The game essentially involves figuring out how to fold proteins in three dimensions. This is an extremely difficult problem, even for computers. The Foldit game is a way of harnessing the brain power of video game players to help find solutions.
The game has been available for a while, and now (apparently for the first time) has been used to solve a specific puzzle of protein structure. Scientists have been trying for 10 years to solve the structure of retroviral protease, a protein-cutting enzyme found in HIV-like viruses. So, they put the problem to Foldit gamers, and they solved the structure in one week.
The reason these types of problems are so difficult is that the number of possible ways in which a large protein can fold is staggering large. This is, fact, called an NP-problem (non-deterministic polynomial). The classic example of this is the traveling salesman who wishes to map the minimum route to a set of cities he wishes to visit. Such problems are impossible to solve by computational brute force (the number of possibilities to check quickly becomes greater than the number of atoms in the universe), and there is no mathematical way to derive the answer quickly.
Continue Reading »
 In 2003 the largest ever international cooperative scientific project was completed, at a cost of about $1 billion – the mapping of the human genome. This came with much fanfare, with the media hyping all the medical benefits that would soon flow. Of course, basic science progress often precedes clinical applications by decades, so the hype was not necessarily wrong, just premature. But it was an immediate boon to research, and those benefits are being felt today.
In 2003 the largest ever international cooperative scientific project was completed, at a cost of about $1 billion – the mapping of the human genome. This came with much fanfare, with the media hyping all the medical benefits that would soon flow. Of course, basic science progress often precedes clinical applications by decades, so the hype was not necessarily wrong, just premature. But it was an immediate boon to research, and those benefits are being felt today.
 I wasn’t planning on writing about artificial intelligence (AI) two days in a row, but I also can’t plan the news. I also couldn’t pass up this item – London-based AI company, DeepMind, has
I wasn’t planning on writing about artificial intelligence (AI) two days in a row, but I also can’t plan the news. I also couldn’t pass up this item – London-based AI company, DeepMind, has 




