Jul 23 2021
AI Advances Mapping of Human Proteome
 In 2003 the largest ever international cooperative scientific project was completed, at a cost of about $1 billion – the mapping of the human genome. This came with much fanfare, with the media hyping all the medical benefits that would soon flow. Of course, basic science progress often precedes clinical applications by decades, so the hype was not necessarily wrong, just premature. But it was an immediate boon to research, and those benefits are being felt today.
In 2003 the largest ever international cooperative scientific project was completed, at a cost of about $1 billion – the mapping of the human genome. This came with much fanfare, with the media hyping all the medical benefits that would soon flow. Of course, basic science progress often precedes clinical applications by decades, so the hype was not necessarily wrong, just premature. But it was an immediate boon to research, and those benefits are being felt today.
Perhaps the next big mapping project in biology is the human proteome, the characterization of every human protein. (I’ll also give a nod to the connectome project, the mapping of every connection in the human brain, but that will likely take much longer.) A new study published in Nature announces a significant leap forward in mapping the human proteome, using artificial intelligence (AI), specifically AlphaFold2 developed by DeepMind. To understand what they accomplished, however, we need to go over some basic concepts and terminology.
A gene is essentially a code for a sequence of amino acids, which make up proteins. So if we have mapped the entire sequence of bases (of which there are four – GATC) in a gene, we know the sequence of amino acids in the protein it codes for. So then, you might ask, if we have already mapped all the human genes, why is that not the same thing as a map of all the human proteins? This is because a protein is more than just a sequence of amino acids. A short chain of 2 or more amino acids is called a peptide, and a long chain is therefore a polypeptide. But we still don’t have a protein. A protein is a polypeptide that folds itself into a specific three-dimensional structure. It is that three-dimensional structure which determined the function and properties of the protein.
In order to go from the genome to the proteome, all we need to know is how to determine exactly how a polypeptide will fold itself based upon the sequence of amino acids. This, in turn, is determined by the chemical properties of the amino acids, and we have figured out the basics of how they affect folding – which ones create a “kink” in the chain, which ones will form into sheets, etc.. But this is still a long way away from predicting how a polypeptide will fold itself into a final protein. There are so many possibilities, in fact, that the “protein folding problem” defies mathematical prediction. Even the most powerful computers, using brute force, cannot solve the protein folding problem (maybe quantum computers will one day).
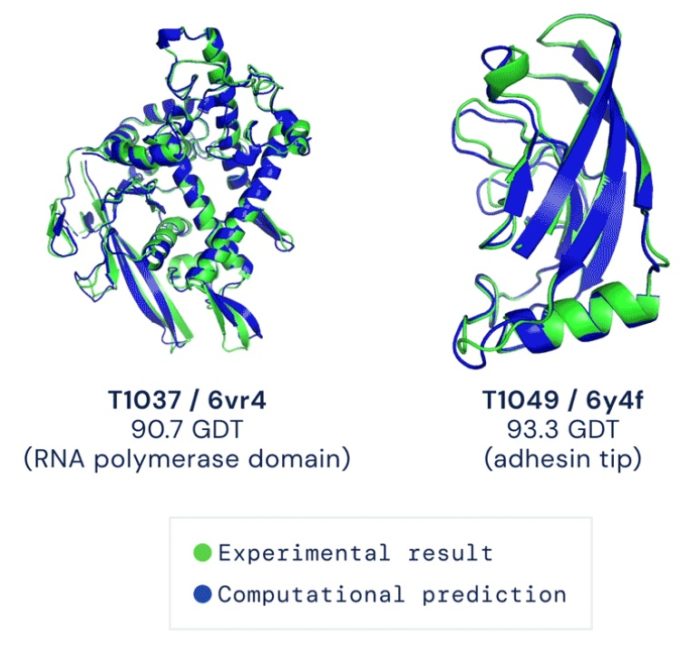
AI, however, has provided the solution. Using deep learning and other AI techniques programmers have partially solved the protein folding problem. The AI software can make high probability predictions about how a polypeptide is likely to fold, into what is known as its “native stable structure.” Where does the current study take us?
“After decades of effort, 17% of the total residues in human protein sequences are covered by an experimentally-determined structure1. Here we dramatically expand structural coverage by applying the state-of-the-art machine learning method, AlphaFold2, at scale to almost the entire human proteome (98.5% of human proteins). The resulting dataset covers 58% of residues with a confident prediction, of which a subset (36% of all residues) have very high confidence. “
A “residue” is simply the part of an amino acid that remains when it is chemically bound to another amino acid in a protein. Therefore, we have experimentally confirmed the position of 17% of amino acid residues in human proteins. The AlphaFold2 project now adds confident predictions from AI to the total number, bringing the total up to 58% (with 36% being highly confident). That is a dramatic increase, and something that we could not have accomplished experimentally in this much time.
This is not the same thing as saying we know the structure of 58% of human proteins, because the database covers portions of 98.5% of human proteins. Also, the 58% figure may be better than it sounds, because there are parts of proteins that are more biologically important that other parts, like the binding region (the business end) of a protein, for example. DeepMind is also making their dataset publicly available, so any researcher can avail themselves of this resource.
Don’t expect medicine to be transformed tomorrow, but I don’t want to undersell the value of this resource either. Knowing the three-dimensional structure of important bits of proteins is extremely valuable to research. It could allow, for example, for computer modeling of the effect of drugs on specific proteins. If you want to design a drug that will bind to and block (or activate) a protein receptor, now you can model that in a computer, because you have the structure of the receptor. This can help researchers understand aspects of cell function, which will help them further understand disease states. How does a genetic mutation alter a protein’s structure and what effect does that have on its function?
Essentially, this database facilitates moving some aspects of biological and medical research from animals and humans into computers (in silico). Computer-based research is much faster and cheaper than dealing with cell cultures or whole animals, and may bypasses ethical complications when they are relevant. The goal is always, before we get to the level of human research, to know as much as possible about risk vs benefit, and mapping proteins will help us do that. We want to use human research simply to confirm high probability predictions made with pre-clinical research. This includes in-vitro and animal research, and increasingly in-silico research.
Innovations which improve research itself have only indirect benefits, and therefore there can be a long lag between the innovation and the clinic – but the impact can eventually become immense. Mapping the proteome will transform the practice of medicine in many ways, but it will be largely behind the scenes, and we will see those benefits down the road. This advance also further shows the potential for narrow AI, which is clearly a powerful tool that is becoming increasingly useful. AI is transforming our world, and it’s just getting started.






