Mar
06
2025
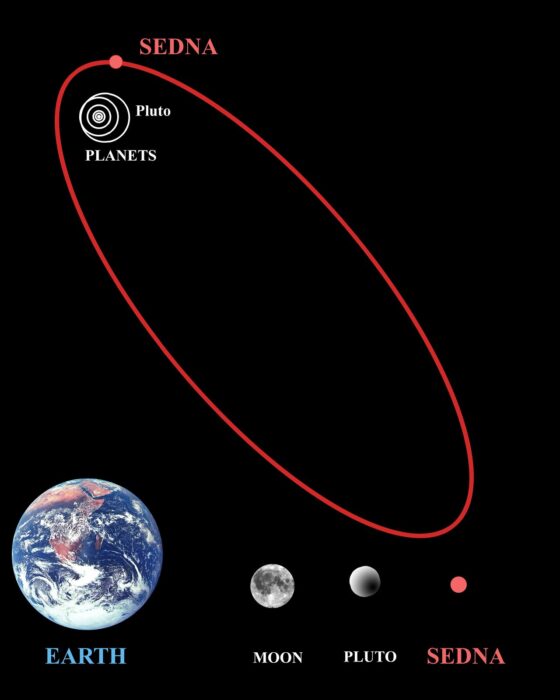
In 2006 (yes, it was that long ago – yikes) the International Astronomical Union (IAU) officially adopted the definition of dwarf planet – they are large enough for their gravity to pull themselves into a sphere, they orbit the sun and not another larger body, but they don’t gravitationally dominate their orbit. That last criterion is what separates planets (which do dominate their orbit) from dwarf planets. Famously, this causes Pluto to be “downgraded” from a planet to a dwarf planet. Four other objects also met criteria for dwarf planet – Ceres in the asteroid belt, and three Kuiper belt objects, Makemake, Haumea, and Eris.
The new designation of dwarf planet came soon after the discovery of Sedna, a trans-Neptunian object that could meet the old definition of planet. It was, in fact, often reported at the time as the discovery of a 10th planet. But astronomers feared that there were dozens or even hundreds of similar trans-Neptunian objects, and they thought it was messy to have so many planets in our solar system. That is why they came up with the whole idea of dwarf planets. Pluto was just caught in the crossfire – in order to keep Sedna and its ilk from being planets, Pluto had to be demoted as well. As a sort-of consolation, dwarf planets that were also trans-Neptunian objects were named “plutoids”. All dwarf planets are plutoids, except Ceres, which is in the asteroid belt between Mars and Jupiter.
So here we are, two decades later, and I can’t help wondering – where are all the dwarf planets? Where are all the trans-Neptunian objects that astronomers feared would have to be classified as planets that the dwarf planet category was specifically created for? I really thought that by now we would have a dozen or more official dwarf planets. What’s happening? As far as I can tell there are two reasons we are still stuck with only the original five dwarf planets.
Continue Reading »
Mar
03
2025
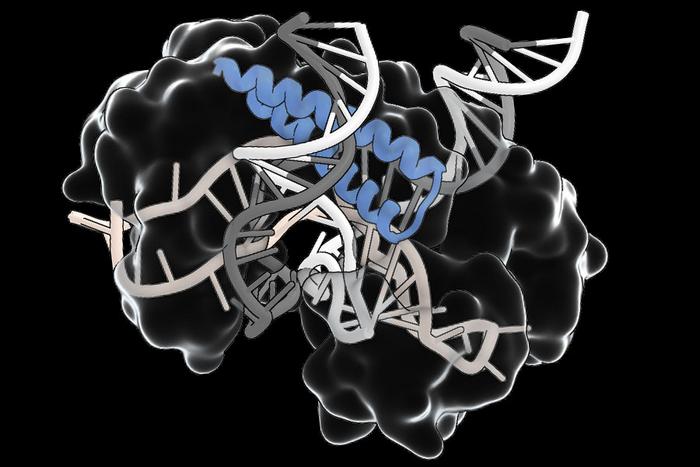 Remember CRISPR (clustered regularly interspaced short palindromic repeats) – that new gene-editing system which is faster and cheaper than anything that came before it? CRISPR is derived from bacterial systems which uses guide RNA to target a specific sequence on a DNA strand. It is coupled with a Cas (CRISPR Associated) protein which can do things like cleave the DNA at the targeted location. We are really just at the beginning of exploring the potential of this new system, in both research and therapeutics.
Remember CRISPR (clustered regularly interspaced short palindromic repeats) – that new gene-editing system which is faster and cheaper than anything that came before it? CRISPR is derived from bacterial systems which uses guide RNA to target a specific sequence on a DNA strand. It is coupled with a Cas (CRISPR Associated) protein which can do things like cleave the DNA at the targeted location. We are really just at the beginning of exploring the potential of this new system, in both research and therapeutics.
Well – we may already have something better than CSRISP: TIGR-Tas. This is also an RNA-based system for targeting specific sequences of DNA and delivering a TIGR associated protein to perform a specific function. TIGR (Tandem Interspaced Guide RNA) may have some useful advantages of CRISPR.
As presented in a new paper, TIGR is actually a family of gene editing systems. It was discovered not by happy accident, but by specifically looking for it. As the paper details: “through iterative structural and sequence homology-based mining starting with a guide RNA-interaction domain of Cas9”. This means they started with Cas9 and then trolled through the vast database of phage and parasitic bacteria for similar sequences. They found what they were looking for – another family of RNA-guided gene editing systems.
Like CRISPR, TIGR is an RNA guided system, and has a modular structure. Different Tas proteins can be coupled with the TIGR to perform different actions at the targeted site. But there are several potential advantages for TIGR over CRISPR. Like CRISPR it is RNA guided, but TIGR uses both strands of the DNA to find its target sequence. This “dual guided” approach may lead to fewer off-target errors. While CRISPR works very well, there is a trade-off in CRISPR systems between speed and precision. The faster it works, the greater the number of off-target actions – like cleaving the DNA in the wrong place. The hope is that TIGR will make fewer off-target mistakes because of better targeting.
Continue Reading »


 Remember
Remember 




