Jul 05 2022
Soft CRISPR For Genetic Diseases
 The technology of CRISPR (clustered regularly interspaced short palindromic repeat) continues to advance as a rapid pace. A recent study, Cas9/Nickase-induced allelic conversion by homologous chromosome-templated repair in Drosophila somatic cells, provides the potential for a new method of treating certain kinds of genetic diseases. This approach also appears to be safer, with fewer off-site genetic changes, but still has to be tested in humans.
The technology of CRISPR (clustered regularly interspaced short palindromic repeat) continues to advance as a rapid pace. A recent study, Cas9/Nickase-induced allelic conversion by homologous chromosome-templated repair in Drosophila somatic cells, provides the potential for a new method of treating certain kinds of genetic diseases. This approach also appears to be safer, with fewer off-site genetic changes, but still has to be tested in humans.
CRISPR works by pairing it with a guide RNA (gRNA) that targets a specific sequence in the DNA, and with a Cas9 endonuclease which will cleave the DNA at the target site. The normal DNA repair machinery will then fix the cut, but there are two basic pathways for this to happen. The first pathway (nonhomologous end joining – NHEJ) blindly reconnects the ends together, creating the potential for the introduction of random mutations at the repair site. This is considered an error-prone repair pathway. The other pathway is homology-directed repair (HDR), which uses the other copy of the DNA as a template, and is therefore much less prone to error. Remember, every cell has two of every somatic chromosome, one from each parent, and therefore two copies of every gene.
The researchers in the current study wanted to know if the HDR pathway could be exploited to not only repair the cleaved DNA but also to make the repaired copy of the gene look like the other copy, the “homologous” gene. Some genetic conditions or traits are dominant and others recessive. A dominant trait will manifest if an organism has only one copy of that trait, which a recessive trait requires that both copies of the gene have the trait. The classic example is eye color (this is an oversimplifcation, but demonstrates the point). Brown eye color is the dominant allele (gene version), while blue is recessive. A person with one brown and one blue allele will have brown eyes (dominant). You need two blue alleles to have blue eyes (recessive).
Therefore, a dominant genetic disease requires only one of the two alleles to have the genetic mutation in order to manifest the disease. While it’s possible to get a double dose of a dominant genetic disease, one from each parent, it’s statistically unlikely. Most people with a dominant genetic disease will have only one copy of the disease-causing allele. That means they also have a healthy allele on the other chromosome. What if, the researchers asked, we can exploit the HDR repair pathway to convert a disease-causing allele into a healthy allele? You just need to target the mutation on the dominant disease allele that results in the disease-causing protein, and then use the HDR pathway to repair it using the healthy allele as the homologous template.
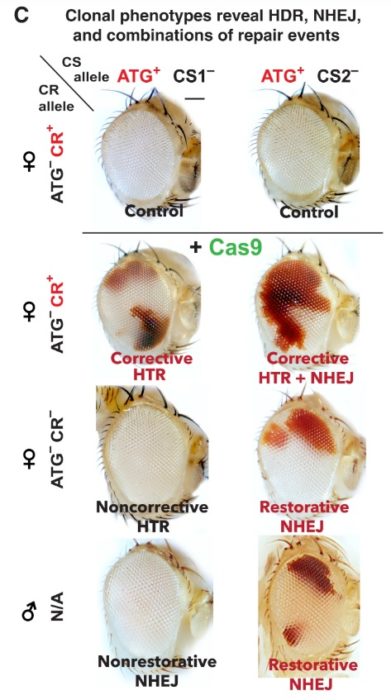
There is one other variable here – when you use CRISPR to make the original cut, do you cut both strands of the DNA or just one strand? Cas9 creates a double-stranded cut. But you can replace Cas9 with an enzyme that will cause only a single-stranded cut, a so-called “nick”. This enzyme is appropriately called a “nickase”. The researchers used Drosophila, a fruit fly, with a dominant mutation that causes their normal reed eyes to become white. This is a common genetic research subject because the researchers can easily see which gene a Drosophila has just by looking at their eyes.
Using the Cas9 double-stranded breaks and the HDR repair, the researchers found the resulting Drosophila had patches of red in the otherwise white eyes. So the homologous repair was working to some extent (20-30%), with some of the cells reverting to the wild-type red trait. Then they tested it with the nickase making only a single strand cut, and found that the HDR repair was more efficient (40-65%). The Drosophila eyes became mostly red, similar to wild-type flies without the white-eye mutation. The nickase was actually more efficient than Cas9 at making the homologous repair.
Also, the single-stranded cuts were much safer, making far fewer errors or off-target changes than the Cas9 double-stranded cuts. So the nickase system was both more efficient and less error-prone than Cas9. This was expected as this was already known, that single strand cuts are safer than double-strand cuts.
Actual clinical applications of this technology is years away, and other tweaks of CRISPR may emerge in the meantime. The next step is to test this system for safety and basic functionality in humans. Then it can be tested in actual disease states. If the technology works this can be a powerful treatment option for dominant genetic diseases. How much will depend on the nature of the specific disease. A 40-65% reduction in the number of cells with the dominant mutation will very likely have a beneficial effect in almost any disease, but in some more than others. Most tissues and organs have built-in redundancy and functional reserve. Your kidneys, for example, can work just fine at 50% capacity.
There is likely to be a steady stream of these kinds of advances in the basic CRISPR technology. It is an extremely useful platform with lots of components that can be altered or swapped out, making it customizable for specific applications. There is always a frustrating delay from the basic science to clinical applications, but we are already seeing some clinical applications using CRISPR, and they are likely to continue.






